Written by Michael J. MacCoss, Department of Genome Sciences, University of Washington
Background: IDCalc was developed to provide a simple way to view the expected isotope distribution for biological molecules measured by mass spectrometry. The program also includes the the ability to consider enriched stable isotopes.
Input: Input is either as a peptide sequence or an elemental composition through the graphical user interface.
Method: The algorithm for predicting the isotope distribution is based on the approach reported by Kubinyi, Analytica Chimica Acta, 247 (1991) 107-119. The C, H, O, and N isotope abundances are specific to biological molecules (Dwight E. Matthews personal communication). All other elemental isotope abundances were obtained from Biemann, Mass Spectrometry, Organic Chemical Applications, McGraw-Hill New York, 1962, p.59
Availability: IDCalc is free for non-profit use. Source code is available upon request. IDCalc is academic software and is use at your own risk.
Platform: IDCalc has been tested using Microsoft Windows XP and Vista.
Note: You might need to install the latest version of the Visual Basic 6 Run Time first before the executable will run on your computer. Go here to download the latest VB6 Run Time libraries from Microsoft. Also, on Windows 7 I have found that you have to register the COMDLG32.OCX library. You can do this by running "regsvr32 C:\SWTOOLS\Apps\MMCfto\System32\Redist\MS\System\COMDLG32.OCX"
Download: Right mouse click here and select "Save link as ..."
Screenshot:
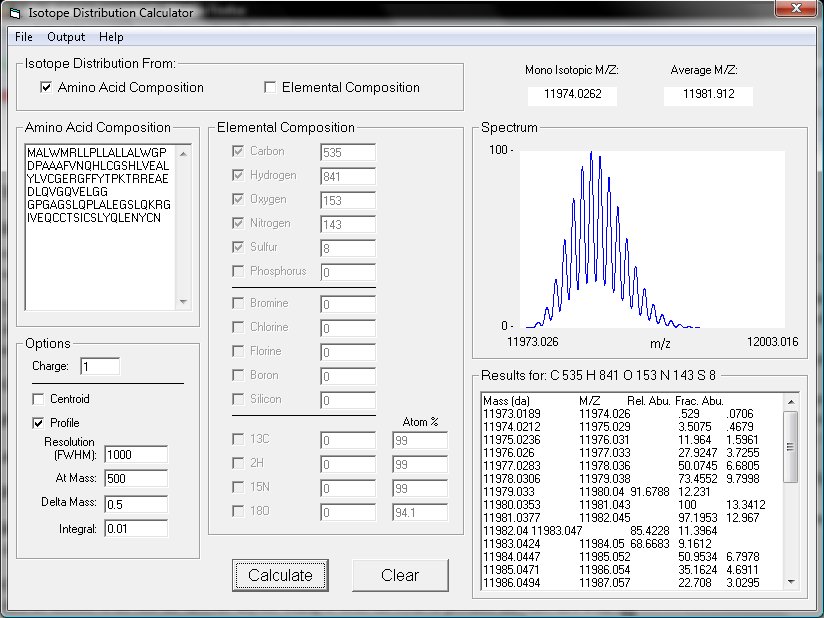
IDCalc is Copyright © 2008 University of Washington. All rights
reserved. Written by Michael J. MacCoss, in the Department of Genome
Sciences at the University
of Washington
MacCoss Lab
Department of Genome Sciences
University of Washington